Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Ekaterina Bazhenova, Irina Kiseleva*, Isakova-Sivak and Tatiana Kotomina
Received: March 26, 2018; Published: April 06, 2018
*Corresponding author: Irina Kiseleva, Department of Virology, IEM, 12 Acad. Pavlov Street, Saint Petersburg, 197376, Russia
DOI: 10.26717/BJSTR.2018.03.000925
Avian influenza remains a major pandemic threat. In response to this threat, the objective of our work was the development of live attenuated reassortant influenza vaccine (LAIV) against recent isolates of potentially pandemic H7N9 influenza viruses which may cause serious and fatal disease in humans. Two H7N9 LAIV candidates were generated by using two different approaches: classical reassortment in eggs and reverse genetics. The collected data indicate that regardless of the approach both H7N9 LAIV candidates acquired attenuated properties - temperature sensitivity and cold-adaptation.
Keywords: Pandemic Influenza; Avian Influenza; Live Attenuated Influenza Vaccine; Potentially Pandemic Vaccine
Abbreviations: LAIV: Live Attenuated Reassortant Influenza Vaccine; CA: Cold-Adapted; HPAIV : Highly Pathogenic Avian Influenza Virus; TS: Temperature Sensitive; WHO: World Health Organization; LPAIV: Low Pathogenic Avian Influenza Virus; RG: Reverse Genetics; SPF: Specific Pathogen-Free; WT: Wild Type
Avian influenza remains a major pandemic threat. Influenza A/ H7N9 virus is a potentially pandemic subtype to which the majority of human population is immunologically naïve. Vaccination is an essential public health tool to control influenza. In the event of a pandemic, the use of live attenuated cold-adapted (ca), temperature sensitive (ts) reassortant influenza vaccine (LAIV) may have a number of advantages over inactivated vaccine; LAIVs that are currently licensed in the Russian Federation, the United States and Europe appear to be safe and efficacious and provide longer lasting and more cross-reactive immune responses compared to inactivated vaccines [1-4]. To ensure optimal vaccine efficacy against currently circulating seasonal influenza strains, the antigenic composition of the vaccines is revised twice a year by the World Health Organization (WHO) [5]. In recent years WHO recommendations also include potentially pandemic viruses. In 2017 WHO recommended developing candidate vaccine viruses from two new potentially pandemic A/H7N9 viruses, A/Guangdong/17SF003/2016, A/ GD (highly pathogenic avian influenza virus, HPAIV) and A/Hong Kong/125/2017, A/HK (low pathogenic avian influenza virus, LPAIV) [6]. The objective of this work was to generate new LAIV candidates against currently circulating H7N9 HPAIV and LPAIV with pandemic potential using two methodological approaches, classical reassortment in eggs and reverse genetics (RG).
a. Master donor virus for Russian LAIV, ca/ts A/ Leningrad/134/17/57 (H2N2), Len-MDV (property of IEM).
b. A/HK LPAIV, (CDC, USA).
c. Plasmids encoding HA (polybasic cleavage site deleted) and NA genes of A/GD HPAIV (NIBSC, UK).
d. Plasmids encoding six internal genes of Len-MDV [7].
Classical Reassortment Procedure: Len-MDV and A/ HK LPAIV were co-infected in specific pathogen-free (SPF) embryonated chicken eggs, CE (VALIO Biomedia GmbH, Germany) as described previously [8].
RG-Generation of LAIV Candidate: 6:2 LAIV reassortant virus (HA and NA genes of A/GD HPAIV and 6 remaining genes of Len- MDV) was rescued by electroporation of Vero cells and amplified in SPF eggs. Mutagenesis of A/GD NA plasmid was performed using overlap PCR strategy.
The LAIV consists of attenuated reassortant viruses incorporating the key immunizing antigens and antigenic determinants (HA and NA) of circulating viruses on a backbone of the remaining six internal protein genes derived from the attenuated MDV (6:2 genome composition) [2]. Generation of 6:2 vaccine reassortant by replacement of internal genes of wild type (WT) virus with the appropriate internal genes of MDV is a routine method of attenuation of WT influenza viruses. Internal viral proteins of MDV provide attenuation of vaccine reassortant [9] ensuring stability of its genetic and phenotypic characteristics and safety when given to susceptible individuals. Surface glycoprotein, HA and NA of the WT virus provide the targets for an adequate immune response in vaccinated host.
A/HK-based LAIV candidate, A/17/HK was prepared through the classical genetic reassortment in SPF CE. In brief, seven rounds of selective propagation were performed, three of which were at low temperature of 26°C. A/17/HK inherited HA and NA genes from WT parental virus A/HK and six internal genes from Len- MDV. The attenuated phenotype of the A/17/HK vaccine strain was confirmed by full-genome sequencing and by virological methods (determination of range of temperature sensitivity and cold adaptation during reproduction in CE). The results showed that the A/17/HK possessed high reproductive activity in CE had a temperature-sensitive and cold-adapted phenotype (Table 1).
Table 1: Reproductive Capacity of H7N9 LAIV Candidates and MDV at Different Temperatures of Incubation in Embryonated Chicken Eggs.
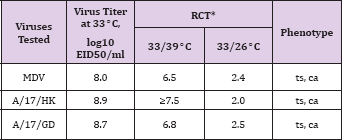
Note: *Reproductive capacity of influenza viruses at different temperatures (ts and ca phenotype) was assessed by titration in eggs at the permissive (33 °C), elevated (39 °C) and low (26 °C) temperatures of incubation and expressed as RCT26 or 39 = log10 EID50/ml at 33°C-log10 EID50/mI at 26 or 39°C. Eggs were incubated for 2 days at 33°C and 39°C and for 6 days at 26°C.
Eight mutations in internal genes of Len-MDV which play an essential role in attenuation of Len-MDV and Len-MDV-based LAIV candidates have been identified and described in [7,10]. A/17/ HK was genetically stable, was identical to the parental WT A/HK virus in antigenic properties and contained in its internal genes all attenuating mutations of MDV: PB2 (V478L), PB1 (K265N, V591I), PA (L28P, V341L), Ml (I15V, F144L) and NS2 (M100I).
In Russian Federation the manufacture of influenza vaccines is based on viral propagation in CE. To lower pathogenicity of HPAIV to CE and to reduce a risk of manipulations of personnel with this virus, we generated cold-adapted A/GD-based LAIV candidate (A/17/GD) on the base of reverse genetically derived H7N9 strain. A/17/GD LAIV candidate displayed classical vaccine genome 6:2 composition - six internal genes of MDV origin and two surface genes were derived from WT virus. The HA protein of the A/17/ GD LAIV candidate had polybasic cleavage site removed, thus making the virus low pathogenic. Neuraminidase of the original H7N9 HPAIV strain had K292 residue, which is the marker for virus resistance to neuraminidase inhibitor. Therefore, NA mutagenesis was performed and the A/17/GD LAIV candidate contains K292R mutation, which renders the viruses sensitive to the NA inhibitor Oseltamivir.
Full-genome sequencing of A/17/GD LAIV candidate confirmed the presence of all eight attenuating mutations specific for MDV. The A/17/GD displayed pronounced ts/ca phenotype (Table 1). Our results emphasized that both H7N9 LAIV candidates have the potential to be used as pandemic vaccines. They displayed pronounced ts/ca phenotype which is comparable with attenuated MDV and were genetically stable (retained all attenuating mutations known for MDV). However, since the A/HK parental virus represents the cluster that caused the majority of human cases in the fifth H7N9 wave [6], and due to the classical reassortment nature, the A/17/HK LAIV reassortant is the preferred candidate for conducting Phase I clinical trial in volunteers.
We thank WHO for the financial support of our work.


