Impact Factor : 0.548
- NLM ID: 101723284
- OCoLC: 999826537
- LCCN: 2017202541
Shreya Shah*1 , Shloka Shloka2, Zachary Maurer2, Chelsea Sidrane2 and Mehul Shah1
Received: April 12, 2018; Published: April 24, 2018
Corresponding author: Shreya Shah, Drashit Netralaya, Chakalia Road, Dahod-389151, Gujarat, India
DOI: 10.26717/BJSTR.2018.04.000995
Objective: To develop an automated system to classify extra ocular diseases
Study Design: Retrospective cohort
Method: The entire dataset consists of about 7,244 labelled images of patients from Drashti Netralaya Eye Hospital in Gujarat, India. Five diseases were selected for classification: Corneal scars, Dermoid Cyst, Strabismus, Ptosis, and Ocular Surface Disease. Histogram of Oriented Gradient feature descriptors were utilized with Support Vector Machines and Logistic Regression. Modern Neural Network architectures were also applied. Bottleneck CNN and Logistic Regression (Balanced) both performed well according to different error measurements. This work outlines the development of a classifier for extra ocular conditions that uses natural, noisy images of faces taken with point-and-shoot cameras.
Outcome Measures: Accuracy of diagnosis
Result: The Bottleneck CNN achieved the highest test accuracy of 77% but Logistic Regression had the highest true positive rate averaged across all classes. We have found accuracy of 92% for strabismus.
Conclusion: Thus, it is possible to develop a classifier from images for a variety of eye diseases.
Keywords: Artificial Intelligence; Machine Learning; Anterior Segment Classifier
This work outlines the development of a classifier for extra ocular conditions that uses natural, noisy images of faces taken with point-and-shoot cameras. Extra ocular conditions are disorders that affect the outside of the eye, as opposed to intra ocular diseases which affect internal components such as the retina. Ophthalmologists can diagnose many extra ocular conditions without examining the internals of the eye. Thus, we believe that these external images will provide a sufficiently rich data set for making predictions.
This study was approval of ethical committee obtained for this study. This is retrospective cohort study. We enrolled extra ocular images from medical records by automated camera. The entire dataset consists of about 7,244 labelled images of patients from Drashti Netralaya Eye Hospital in Gujarat, India. Five diseases were selected for classification: Dermoid Cyst, Strabismus, Ptosis, and Ocular Surface Disease and Corneal scars (Table 1). The images were pre-processed before they were passed to the classifier. Facial landmark features in each image were located, and then the images were cropped to 150x300, and transformed so that the eyes were center and level in the frame. This processing was done using Dlib [1] and Open Face [2]. The last step was to normalize the image histogram to increase the contrast of the image using the Pillow image processing library [3]. After pre-processing, the dataset consisted of 5,058 labelled images. Eighty percent of the images were used for training and twenty percent were used for testing. A table of image distributions are below (Table 1).
Histogram of Oriented Gradients, or HOG features, were chosen for this project because they have proven very successful for object recognition. In addition, HOG features are resistant to changes in exposure, which was an asset since the exposure of different images in the dataset varied widely. To avoid our models learning from irrelevant artefacts in the original 150x300 image, we also generated another set of features called 'cropped HOG'. To generate these, we extracted HOG features from tightly cropped images of each eye (cropped to 96x96) and concatenated the feature vectors together. The ordering of feature extraction and then concatenation was intended to avoid the introduction of new image artefacts from extracting HOG features from the concatenation of two discontinuous regions of a photograph. For reasons that will be elaborated on later, Principle Components Analysis (PCA) was also used to reduce the dimensionality of the feature space. Lastly, we confirmed that HOG features performed better than the mean brightness of the image, which was chosen as a baseline comparison and yielded classification only slightly better than chance (Figure 1).
Three different models, SVM with a Linear kernel, SVM with an RBF kernel and Logistic Regression were used because they reflect previous work on this subject, and these classifiers are easy to use with the continuous-valued HOG features. The cost function, loss, and kernel are given as follows for the SVM with a Linear Kernel:
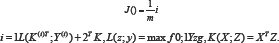
For the SVMP with a Radial Basis function (RBF), the kernel is given by 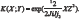
For Logistic Regression the cost function is given by 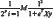
For each of these, a balanced version of the model was also used, which automatically adjusts weights inversely proportional to class frequencies in the input data. These models were implemented using Python's Scikit Learn library [4]. In addition, different architectures of a Convolutional Neural Network were developed using Keras and Tensor flow, based on the information found on the Keras blog [5]. The different architectures we used were: Neural Network In addition, different architectures of a Convolutional Neural Network were developed using Keras and TensorFlow, based on the information found on the Keras blog [5]. The different architectures we used were
Simple CNN: This is a very basic neural network with three layers of 32, 32, and 64 units respectively. Each of these layers has ReLu (Rectified Linear Unit) and maxpooling layers (to reduce dimensionally). Two dense layers were added on top and a final unit using sigmoid activation was used for making predictions. The loss we used was binary cross entropy.
Bottleneck CNN: This CNN used the bottleneck features generated by running the images from the dataset once through VGG16 (a 16-layer neural network used by the VGG team in the ILSVRC-2014 competition), instantiated only for the convolutional layers. These features were then used to train a fully connected CNN. The net has a fully connected layer followed by a dropout layer (for regularization) and a final prediction unit with sigmoid-softmax activation for multi-class predictions. This CNN also uses binary cross entropy for loss (categorical cross entropy for multiclass) and accuracy as the training metric. We used RMSprop as the optimizer for binary classification, but this resulted in a sudden drop in accuracy during training for the multiclass case. We tried several other optimizers (stochastic gradient descent, Adam), and Adam turned out to be the most performant.
VGG16 (Topmost Layers): In this model, we fine-tuned the last convolutional block of VGG16 along with the fully connected CNN and froze all the other layers for VGG16 while training. We used Stochastic Gradient Descent as the optimizer and accuracy as the training metric. Before training, we also performed data augmentation on the images using Keras’s Image Data Generator, which generates similar images by rotating, zooming in randomly, translating horizontally or vertically, shearing and randomly horizontally flipping the images.
Metrics the F1 score, which is the harmonic mean of precision and recall, was chosen as a performance measure because it reflects the relevance of a given disease classification to an image. While many diagnostic tools use sensitivity and specificity, because we do not consider a "no disease" classification, individual sensitivity and specificity values have less significance. However, the F1 score is biased measure and is affected by the distribution of samples in each category. For that reason, during cross validation and parameter tuning, we used an additional metric, the Area Under Curve of the Receiver Operating Characteristic curve (AUC-ROC) to encourage the selection of "well-rounded" classifiers.
Linear Classifiers The performance of the selected models was first assessed using a binary classification task to give an estimate of the feasibility of this project, and to predict the situations where a more complicated multi-class model would have difficulty making predictions. For these binary tests, the three disease datasets with the largest number of cropped images were used: Strabismus (str), Ptosis (ptosis), and Ocular Surface Disease (OSD). Initial results using these data demonstrated that SVM models with a Radial Basis Function (RBF) kernel performed the worst. It was suspected that this poor performance was due to a lack of tuned hyperparameters and possibly the high dimensionality of the feature space. The initial results from binary classification are show in the Table 2.
To examine these results, learning curves were created for the top performing models. A subset of these graphs is shown below. These curves plot the test and train error for models trained on increasing training set sizes which were additively generated and separate from the test set. From these graphs, we hypothesized that our classifiers had high variance because training error was very low and test error much higher (Figure 2). Neural Network Classification We ran all three neural nets for classifying Ptosis vs. Strabismus. All three neural nets overfit the training data. During training we saw that Simple CNN achieved around 84% after 500 epochs (the exact accuracy was 84.1% for training, and 79% test accuracy). The top layer CNN using VGG16 had around 92% accuracy during training after 100 epochs. The bottleneck CNN performed the best, with a train accuracy of 98% and a test accuracy of 88%. We ran the Bottleneck CNN for the multiclass case because it ran in under 5 minutes, as opposed to the top layer CNN which took around 18 hours to run 200 epochs.
Following the initial exploratory model development, all selected models were tested for the multiclass case with five diseases: Strabismus, OSD, Ptosis, Corneal Scars, Dermoid Cyst. The best performing classifier with standard HOG features was Logistic Regression (Balanced) (Table 3). Linear Classifiers Drawing on the conclusions from the learning curves derived from the binary classifiers, we sought to reduce the variance of our models by (3) introducing 'cropped-hog' features (which reduces the size of the original image and consequently the number of HOG descriptor values) and (1) applying Principal Components Analysis to our features to reduce their dimensionality. These Logistic Regression (Balanced) using cropped HOG features, and Logistic Regression (Balanced) using HOG features and a PCA reduction to 4,037 principle components compared very similarly to Logistic Regression (Balanced) with standard HOG features. Once the performance of PCA-transformed features was established as reasonable, estimates for the ideal number of principle components were obtained by plotting the fraction of explained variance as a function of the number principal components.
These plots are shown below. The key take-away is that there is minimal variance explained beyond 1000-2000 principal components. Cross Validation and Grid Search To improve upon these results, 5-fold cross validation was performed using exhaustive grid search to tune the hyper paramaters of all multi-class models listed above. AUC-ROC was used as the cross validation scoring metric. This was designed to correct for the imbalance of class examples, as explained in the Metrics section. The best models resulting from evaluating 40-150 candidate tunings for each classifier is listed (Table 4) and (Figure 4) Most notably, there was significant improvement in the performance of the RBF-SVM. However, despite these improvements, Logistic Regression performed more poorly with 1000 or 2000 principal components than with a greater number of features, as previously observed. This is likely due to the loss in predictive signal caused by reducing the number of principal components. As a final tuning step, the best hyperparameters for each model that were derived from cross validation were then applied to models which trained on the full set of uncompressed cropped HOG features (Figure 3). Final Results The best classifiers chosen using model selection are compared in Table 5. The best performing multiclass classifier was the Bottleneck CNN when comparing using the F1 score, the area under ROC curve computed during validation, and the accuracy (Table 5).
Most of existing research on using machine learning techniques to detect eye diseases in images of a person's eye or face are taken with specialized camera equipment or utilize specialized diagnostic data [6,7]. These efforts have generally utilized models like Support Vector Machines (SVM) and neural networks (NNs) to make predictions [6]. Research has also been done on classifying retinal scans/images for particular diseases, such as diabetic retinopathy, which can lead to blindness [8-10]. There are ongoing efforts between Google DeepMind and the NHS to evaluate automated techniques for interpreting more specialized eye scans [11]. There have also been efforts to build a web-based diagnostic system for diabetic retinopathy relying on human experts in order to identify early onset of the disease [10-14]. However, there seem to be comparatively few research studies on diagnosing extra ocular conditions, especially for photos that are not taken with specialized equipment. Most previous work also involves detecting the presence or absence of a single eye condition, as opposed to classifying multiple different eye disorders. Thus, we believe that our work: attempting to build a multi-class extra ocular eye disease classifier by applying modern methods to a heterogeneous image dataset taken with point-and-shoot cameras, will be a welcome extension on existing research.
In this paper, we have demonstrated an iterative tuning effort to produce classifiers that best classify extra ocular diseases from conventional images of people's faces. Suspected high variance It was suspected that initial poor performance of the RBF-SVM was due to a lack of tuned hyperparameters (radius for the RBF was too large) and the large number (45,000) of features. In addition, the learning curves suggested that logistic regression may had high variance. Hyperparameters tuning and use of PCA-reduced features (Table 4) was so successful that the RBF-SVM ranked above Logistic Regression and the Linear SVM among the candidates tried in the parameter search. As a result, cross-validation for the RBF- SVM using the first 2000 principle components of the data were tried, but 1000 principle components yielded a higher AUC score. In contrast, the performance of logistic regression and the linear SVM, according to the F1 score, was significantly reduced from models using full HOG features. These results may be interpreted to mean that the radial basis function initially had too large of a radius and wasn't able to construct a complicated enough decision surface. As the F1 score for the RBF-SVM and the AUC score got respectively worse and better when removing PCA, we are not able to conclude whether there was high variance in this model. It is possible to conclude that there was likely not high variance in logistic regression and the linear SVMs as removing PCA increased the F1 scores and AUC scores for these models.
Through our tuning efforts, absolute gains were made for the CNN and the RBF-SVM models. However, for most other models, tuning exchanged the distribution of errors across different classes, as opposed to producing a classifier with truly better predictive capability for all diseases. Although not picture here, tuning the Logistic Regression model through cross-validation in attempt to pick the optimal number of PCA-reduced components produced a classifier that misclassifies almost all Dermoid Cyst and Corneal scars examples, while the model without PCA tuning (Figure 3) has accuracy 40-50% on those classes. We believe this to attributable to overfitting during cross validation or a loss of signal due to PCA transformation.
Linear classifiers such as logistic Regression (Balanced) perform similarly to modern neural network architectures such as a bottleneck CNN in classifying Strabismus, Ocular Surface Disease, Ptosis, Corneal Scars and Dermoid Cyst from natural, noisy images of faces taken with non-specialized equipment. Use of these two models in combination with HOG features extracted from the region around the eyes yields test accuracy of 75-77%. Better diagnostic accuracy would be required for deployment in a real system, but this proof-of-concept system demonstrates that automatic classification of this type is possible. In the future we hope to be able to improve our results by pursuing several extensions to our work. We could refine pre-processing, specifically the step where we identify the region corresponding to the eye, to utilize more of the original dataset. To improve the accuracy of the existing neural network, we could try obtaining more images for some of the smaller datasets (for example Corneal Scar) and try running VGG16 and the simple CNN for longer iterations. We could also try and reduce the overfitting by using several methods such as dropout, more aggressive data augmentation and using of L1 and L2 regularization. Apart from making changes to the existing infrastructure, we could also obtain more data to classify whether a patient has an eye disease or is healthy (no disease), and we could classify some of the other more common eye diseases that don't necessarily need surgery (like conjunctivitis, Stye etc.). This would make our classifier more useful.


