Introduction
Emergence of multidrug resistance bacteria is a significant health threat globally. Administration of antibiotics against pathogens causes dysbiosis of gut and development of multidrug resistant strains within the gut and thereby spread of resistance through horizontal and vertical transfer of resistant gene [1]. An alternative to reduce the usage of antibiotics is probiotics, the beneficial microorganisms inhabiting gut. Lactic acid bacteria (LAB) is the most exploited probiotic group which is administrated orally as food supplements and as pharmaceuticals. Probiotics are generally regarded as safe due to lack of pathogenicity. The additional health benefit of LABs like tolerance to lactose, reduction in severity of antibiotic associated and infectious diarrhea, antibacterial effects, anticancer and antimutagenic activity antiinflammatory and protective effects in IBD, cholesterol assimilation etc. increases its use in food and pharmaceutical products [2].
One of the major facts that must be considered while using probiotics is the development and transfer of antibiotic resistance. LABs inhabiting the gut are directly exposed to the antibiotics consumed and its misuse and overdose leads to the development of resistance and thereby the transfer of antibiotic resistant gene .The study aims to evaluate the antibiotic susceptibility of a panel of LABs isolated from breast milk, which is the source of bacteria to infants gut and the LABs from infant feces which might be the residents of infant gut.
Materials and Methods
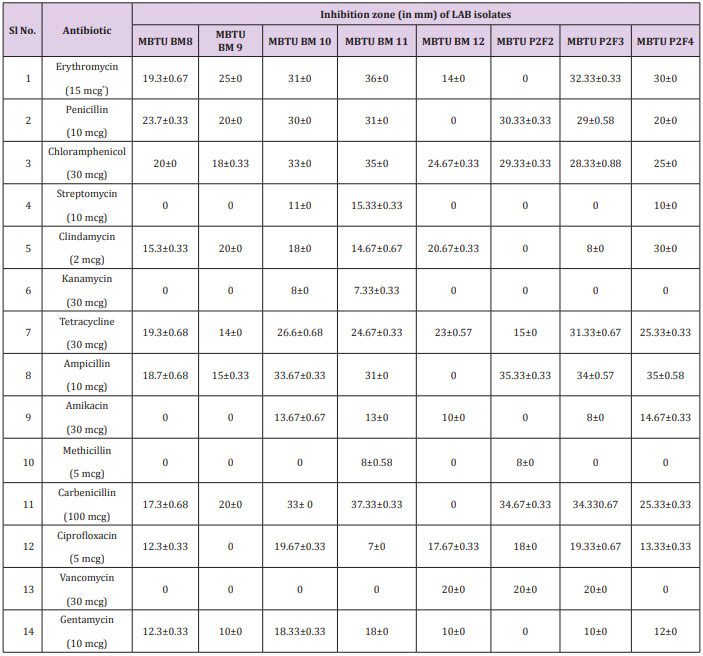
The selected lactic acid bacterial strains were isolated from infant feces and breast milk collected from General Hospital, Kottayam, Kerala, India. Antibiogram of selected bacteria were done by Kirby Baeur disk diffusion. Briefly, isolates were inoculated in MRS broth and incubated, and cell numbers were adjusted to 108 CFU/ml. The cultures were spreaded on MRS agar using sterile cotton swabs. Thirteen commonly used antibiotics were tested which includes erythromycin, penicillin, chloramphenicol, streptomycin, clindamycin, kanamycin, tetracycline, ampicillin, amikacin, methicillin, carbenicillin, ciprofloxacin, vancomycin, and gentamycin. Incubated at 370C for 24 hrs. aerobically. After incubation the zone of inhibition was measured.
Results and Discussion
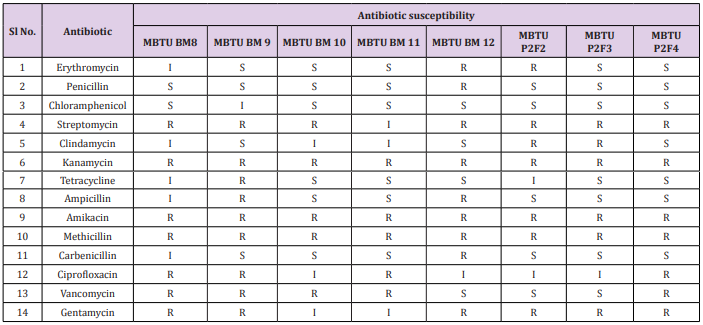
The zone of inhibition was measured after 24 hrs. of incubation. The interpretations were done according to CLSI (2015). The inhibition zone diameter of isolates less than or equal to 14 mm was considered as resistant, zone diameter more than 20 mm as sensitive and zone diameter in between 14 and 20 mm as intermediate. Antibiotic susceptibility profile is extremely different between isolates. Penicillin, carbenicillin and chloramphenicol were effective to most isolates except one isolate as shown in Tables 1 & 2. Previous studies also indicated that LABs were sensitive to β lactams and chloramphenicol [3]. All the isolates showed resistance to amikacin, methicillin, gentamycin and kanamycin. There is an intrinsic resistance to kanamycin, gentamycin, streptomycin and vancomycin by LAB. Resistance to the aminoglycosidic antibiotics kanamycin and streptomycin is due to lack of cytochrome mediated electron transport in LAB which mediates the uptake of antibiotics. The best characterized resistance in LAB is its resistance to vancomycin due to the lack of binding site in peptidoglycan [4]. Interestingly we have got 3 isolates which are vancomycin sensitive. Our results not strictly follow previous studies which indicate these isolates are wild strains that are not more exposed to antibiotics. Commercial LABs are more resistant since they are continuously exposing to different environments and drugs [5] and they act as a reservoir for resistant genes [6].
Table 2: Antibiotic susceptibility profile of LAB isolates, R: Resistant, S: Sensitive, I: Intermediate.
Conclusion
In our study the LAB isolates showed sensitivity towards chloramphenicol penicillin and carbenicillin and complete resistance to methicillin, streptomycin, kanamycin and amikacin. The isolates from the host specific natural sources showed varying sensitivity to most of the antibiotics tested and it also creates a hurdle in the case of combination therapy along with antibiotics. The emergence of multidrug resistance in probiotics is a serious issue as they are dietary supplements and of pharmaceutical importance. Before considering a probiotic the resistance towards antibiotics must be evaluated to recognize them as safe.
Acknowledgement
The authors hereby acknowledge Kerala State Council for Science Technology and Environment (KSCSTE) for the financial support.
References
- Casals pascual C, Vergara A, Vila J (2018) Intestinal microbiota and antibiotic resistance: Perspectives and solutions. Human Microbiome Journal. Elsevier 9: 11-15.
- Kechagia M, Dimitrios Basoulis, Stavroula Konstantopoulou, Dimitra Dimitriadi (2013) Health Benefits of Probiotics: A Review. ISRN Nutrition 2013 p. 1-7
- Shazali, N, Foo HL, Loh TC, Choe DW, Abdul Rahim R, et al. (2014) Prevalence of antibiotic resistance in lactic acid bacteria isolated from the faeces of broiler chicken in Malaysia 6(1): 1–7.
- Lethycia H, Crisley Aparecida Morete, Osmar Roberto DallaSanta, Fernando Bittencourt Luciano, Humberto Maciel França Made, et al. (2017) Methods for the evaluation of antibiotic resistance in Lactobacillus isolated from fermented sausages. 47(8): 1-7.
- Z Erginkaya, EU Turhan, D That (2018) Determination of antibiotic resistance of lactic acid bacteria isolated from traditional Turkish fermented dairy products. Iranian journal of veterinary research 19(1): 53-56.
- Toomey, N, Monaghan A, Fanning S, Bolton D (2009) Transfer of Antibiotic Resistance Marker Genes between Lactic Acid Bacteria in Model Rumen and Plant Environments. 75(10): 3146-3152.
- (2015) In: CLSI Document M100- S22. Clinical Laboratory Standard Institute ‘Performance standards for antimicrobial susceptibility testing: twenty second informational supplement’.
 Case Report
Case Report