Abstract
Background: Every year many organic wastes pollute the environment. One of them is WCW. Biofilm production has been specifically investigated using this waste with high protein and carbohydrate loads.
Aim: whey has become the subject of evaluation in order to obtain different valueadded bio-products or to reduce pollution potential. In this context, whey is thought to be a convenient, cheap and attractive source of biofilm formation by P. aeruginosa.
Method: The same amounts of cells were grown at 37 °C, 0 rpm on incubator for overnight (O/N). Biofilm mass was finally determined as a function of the concentration of this dye based on the absorbance at 570 nm. Swimming and swarming assays was determined using modified methods with WCW as the substrate.
Results: WCW contributed to the formation of cfu/ml, bacterial density and biofilm, while causing a decrease in BFI values. The highest biofilm production occurred at 200 rpm, the highest number of cells; the cell density (live + dead) at OD600 does not support this. Motility plays an important role in biofilm formation and movement in different environmental conditions, colonization, and adhesion of bacteria to surfaces. The lowest swimming was 3 (mm) in agar medium and the highest value was 42.67 (mm) with the addition of WCW. The lowest swarming was carried out in agar medium with 7.66 (mm), while the highest value was found in N.A medium with the addition of 10% WCW 59.33 (mm). In all experimental conditions, an increase of 2.4 times (swimming) and 6.4 times (swarming) was observed after the addition of WCW to the controls.
Discussion: WCW contributed to the formation of cfu/ml, OD600 and biofilm, while causing a decrease in BFI values. As a result, the addition of WCW causes a significant increase in both swarming and swimming.
Keywords: Waste cheese whey; Biofilm; Pseudomonas aeruginosa; Cfu; Biomass; Motility
Introduction
Waste Cheese Whey (WCW) is a liquid side that remains after the cheese making process and represents 95% of the initial milk volume and is considered an important contaminant due to high chemical oxygen demand. Dry matter in whey is up to 7%. These dry ingredients are; lactose, proteins, salts, lipids, lactic acid, citric acid, minerals, vitamins, urea and uric acids. Lactose form 75% of the dries matter in whey. A protein account for 12-14% of this dries matter. This content represents a good opportunity for valuation of whey, an important industrial by-product (for microbial growth) [1-6]. The removal of whey is an important pollution problem especially for the countries connected to the milk economy. Only half of the whey produced each year is converted into useful crops such as food and animal feed and is considered a backwater pollutant [1-3,6]. Microbial biofilm formation involves a variety of surfactant molecules categorized by their chemical composition and microbial origin [1-3,6]. Biofilms provide an environmentally friendly and sustainable alternative in addition to biological production via microbial fermentation from a cheap raw material such as whey [7].
Motility plays an important role in the virulence of P. aeruginosa. This feature plays an important role in the mobilization and colonization of bacteria in different environments, the attachment of bacteria to the surfaces and the formation of biofilm. P. aeruginosa is unusual in that it has three different mobility: swimming in the aquatic environment and in low agar concentrations (0.3 % agar) to flagellum; IV type pilus on solid surfaces mediated swarming and recently observed, swimming over semi-solid (viscous) medium (0.5 to 0.7 % agar). Swimming is generally defined as a dendriticlike colonial appearance and a social phenomenon that typically involves coordinated and rapid movement of bacteria along a semi-solid surface [8-10]. The swarming requires the production of a functional flagellum and rhamnolipid surfactant. The sliding motion is separate from the swimming motion because swimming is the movement required to move on an aqueous, viscous semisolid surface while allowing movement in a liquid medium of relatively low viscosity [11]. The swarming requires the production of a functional flagellum and rhamnolipid surfactant. The sliding motion is separate from the swimming motion because swimming is the movement required to move on an aqueous, viscous semisolid surface while allowing movement in a liquid medium of relatively low viscosity [12-19].
During log-phase growth, P. aeruginosa has a single and polar flagellum that allows for swimming movement. Swarming is a flagellum- dependent community behavior on surfaces. This phenomenon includes many bacteria, including those using polar flagellum during swimming; aggregation is associated with the production of multiple lateral flagella [20]. Pseudomonas aeruginosa is a very rarely resistant nosocomial pathogen. Plants can infect various hosts such as nematodes and mammals. In humans, patients with cystic fibrosis are important opportunistic pathogens in compromised individuals, such as severe burns and impaired immunity [2]. As a result, whey has become the subject of evaluation in order to obtain different value-added bio-products or to reduce pollution potential. In this context, whey is thought to be a convenient, cheap and attractive source of biofilm formation by P. aeruginosa [21].
Materials and Methods
Microorganism
P. aeruginosa (ATCC 27853), obtained from the ATCC and used this study.
Waste Cheese Whey
Waste cheese whey (WCW) was collected from commercial cheese factories in Malatya, Turkey. This waste was filtered for removing crude impurities and then, WCW autoclaved, and then used.
Growth Conditions
Pseudomonas aeruginosa was firstly cultured in Luria- Bertani (LB) broth medium (g l-1); peptone (10), NaCl (10), and yeast extract (5). The final pH values of broth media media were adjusted to 7.0. The same amounts of cells were grown at 37 °C, 0 rpm on incubator for overnight (O/N). 100 μl of overnight cultures (OD600 ~ 0,2-0,3) grown tube filled with 5 ml in 10 ml tubes was inoculated and incubated for 24 h of time. Phosphate-buffered saline (PBS buffer) (gl-1: 8,0 NaCl; 0,2 KCl; 1,44 Na2HPO4; 0,24 KH2PO4 and pH 7,4) and PBS+10 % WCW. These cultures were subsequently incubated on an orbital shaker at 0, 100 rpm, 200 rpm and 37 °C for 24 h.
Biofilm Formation
After the incubation, the supernatant was removed. Biofilm tubes were washed four times with 1 x phosphate buffered saline (PBS) to eliminate any remaining cells. Cells attached to the tubes were then fixed with ethanol (99%) for 15 min room temperature and stained with 1% crystal violet. After staining, excess crystal violet was eliminated with water, and 33 % acetic acid was used to dissolve the remaining dye. Biofilm mass was finally determined as a function of the concentration of this dye based on the absorbance at 570 nm [22- 26]. Biofilm formation was standardized to growth with the biofilm index (BFI), which was calculated. The extent of biofilm formation was determined by applying this formula: “BFI = (AB - CW) ⁄ G in which BFI is the “Biofilm Formation Index was defined as the average percentage of bacteria grown as biofilms [27].”, AB is the OD570 nm of stained attached bacteria and CW is the OD570 nm of stained control tubes containing only bacteria-free medium, G is the OD600 nm of cells growth in suspended culture” [28-29]. OD600 and OD570 were measured using a spectrophotometer directly from tubes. Growth curves were established by plotting the log10 cfu/ml as a function of time. Bacterial growth was determined by measuring the absorbance at 600 nm (OD600) by a spectrophotometer.
Motility
Swimming and swarming assays was determined using modified methods with WCW as the substrate. Swimming. Swimming plates were composed of 0.3 % Nutrient Agar, supplemented with 10 % WCW and sterile phosphate-buffered saline supplemented with 10 % WCW. Swarming. Swarm plates were composed of 0.5 % Bacto Agar and 8 g/L of nutrient broth, supplemented with 10 % WCW and sterile PBS buffer supplemented with 10 % WCW petri dishes. Petri dishes dried overnight at room temperature. Cells were point inoculated with a sterile pipette 6 μl, and the plates were incubated at 25 and 37 °C for 24 hours, respectively. Motility was then assessed qualitatively by examining the circular turbid zone formed by the bacterial cells migrating away from the point of inoculation [10,13,19,20,30-32]. Each value is the average of three independent experiments. Plates were photographed with Nicon Coolpix L320. Each value is the average of three independent experiments.
Results
Biofilm
When we look at the OD570 values the greatest increase was at 2.967 at 200 rpm ventilated runs while the lowest rate was achieved at 0.403 at 0 rpm ventilation conditions. The addition of WCW caused the biofilm formation to decrease at only 100 rpm. The reason for this could not be understood. While this decline is seen in 1.2 times and 100 rpm ventilated conditions, 1.4 times increased at 200 rpm. No significant difference was observed at 0 rpm. Overall, biofilm 10% WCW addition at 0 rpm and 200 rpm resulted in a 7.4-fold increase in biofilm formation and at 0 rpm and 100 rpm resulted in a 3.8-fold increase in biofilm formation (Figure 1). This value is an indication that the contribution of WCW to biofilm formation is high. In another unpublished study, NB the rich media medium, was used. In the end (OD570); 0 rpm 0.066, 100 rpm 0.222, and 0.389 at 200 rpm was obtained (N.A), respectively. This work was done to control.
BFI
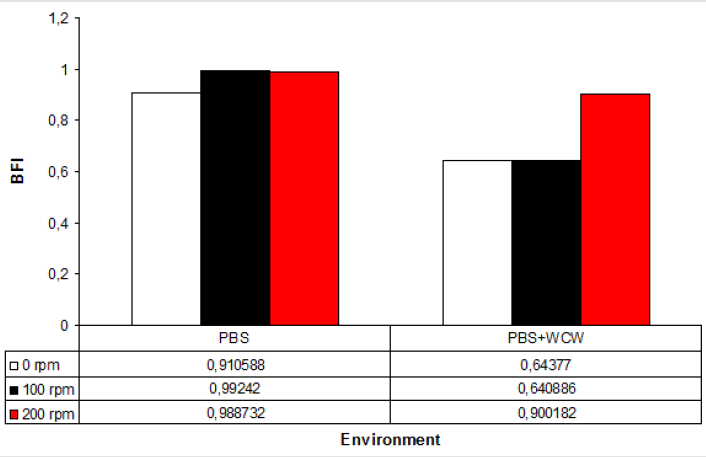
In all experimental conditions, the addition of WCW to controls resulted in a 1.5-fold decrease. The lowest BFI value has 0.64, while the highest value was 0.90 at 200 rpm shaking conditions with addition of WCW. Given the experimental conditions, WCW addition resulted in a reduction of up to 64 % in BFI formation (Figure 2).
CFU
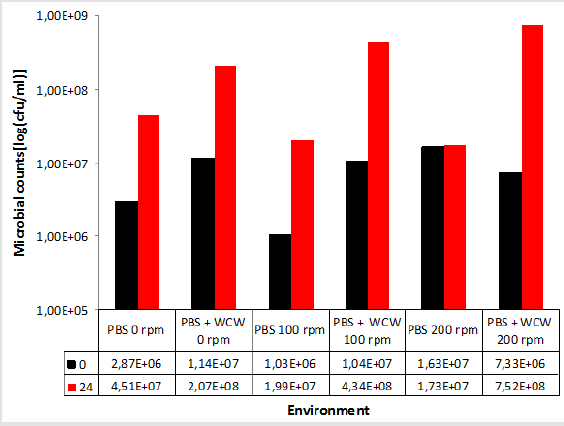
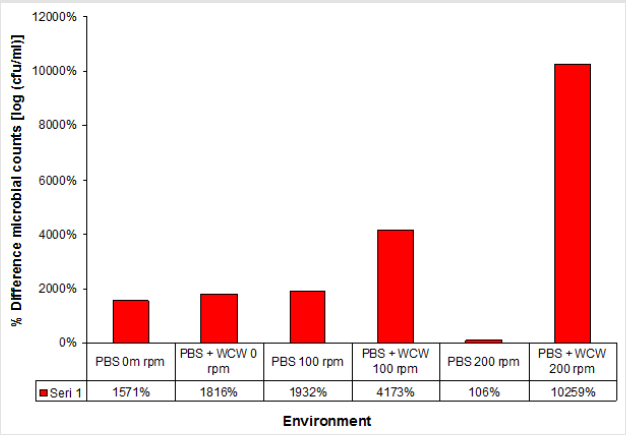
When the PBS medium was not taken into account, the increase in cell count in the WCW-added medium was 7.52x108 at ventilated runs of at most 200 rpm, while the lowest increase occurred at 0 rpm with 2.07x108 non-agitated media. As can be seen, as the ventilation conditions increase, the number of cells increases. However, the difference between 0 rpm and 200 rpm was only 0.3 times (Figure 3). Therefore, ventilation did not make any significant difference when evaluated at 24 hours. When the 24th and the final run were compared, the highest difference was observed at 200 rpm shaking conditions, with the highest difference being 102 times, while the lowest difference was observed at 18 times at 0 rpm with non-shaking conditions. When we look at the difference; the cell count increased by as much as 1.2 times in 0 rpm conditions, 2.2 times in 100 rpm shaking conditions, and 97 times in 200 rpm shaking conditions. The biggest difference was realized at 10259 % at 200 rpm. This was followed by 100 rpm with 4173 % and 0 rpm with 1816 % (Figure 4).
Bacterial density (OD600)
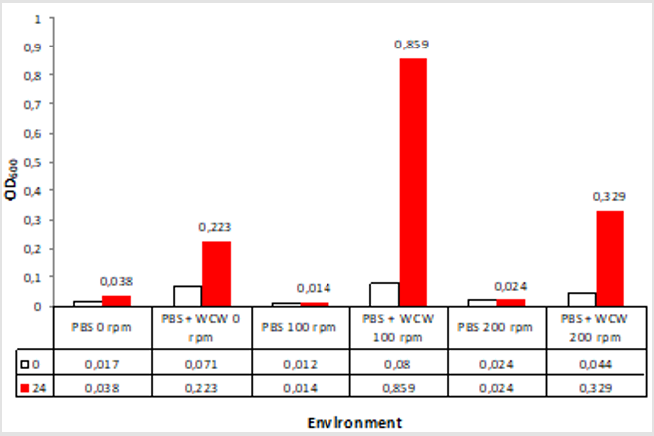
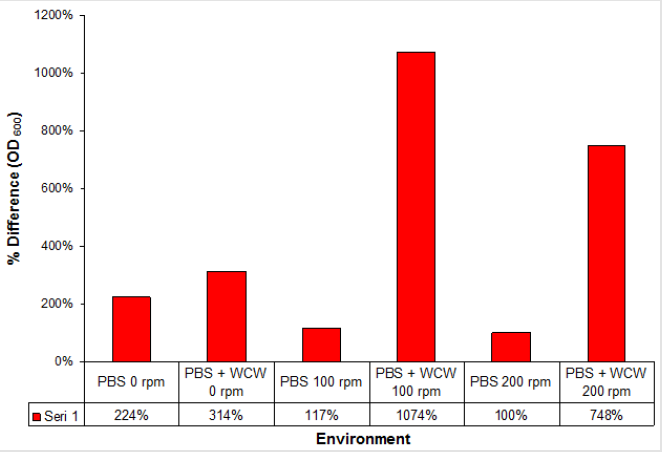
When we look at the OD600 values that we call biomass when adding WCW; again the greatest increase was at 0.859 at 100 rpm ventilated runs while the lowest rate was achieved at 0.233 at 0 rpm ventilation conditions. The difference between 0 rpm and 200 rpm was only 1.5 times. The 24th time that the start and end of the run was compared when the time was compared to 11 times with 100 rpm shaking conditions while the lowest rate was 3.1 times with 0 rpm without agitation. As a different point of view; the highest rate was observed at 9.2 folds with 100 rpm shaking conditions, while the lowest rate was achieved with 1.4 times at 0 rpm non-shaking conditions (Figure 5). When we look at the difference; the largest headlights were followed by 1074 % at 100 rpm followed by 748 % at 200 rpm and 314 % at 0 rpm, which is the 0-rpm condition. Addition of WCW resulted in an increase in biomass values of up to 23-fold when the initial value of 0.038 was considered (Figure 6).
Motility
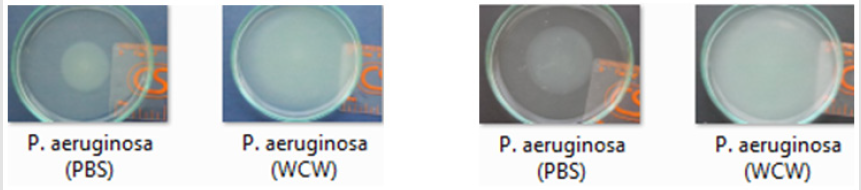
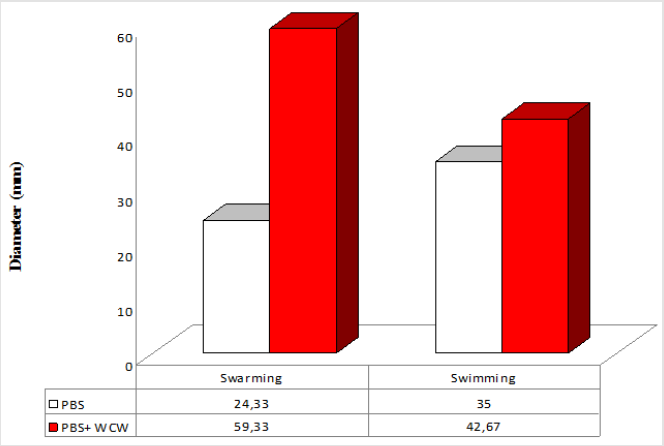
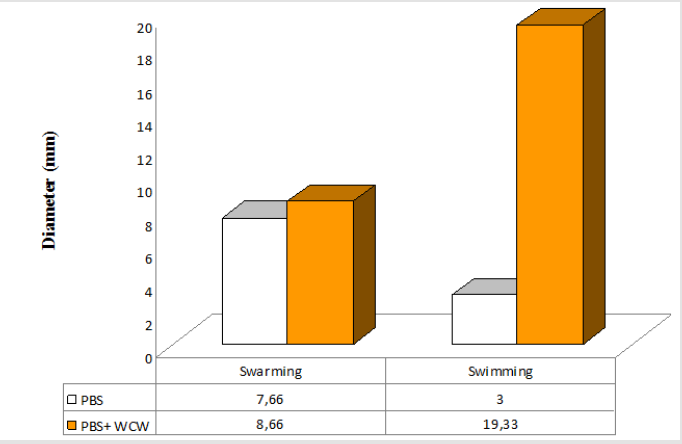
P. aeruginosa is able to swarm on semisolid agar (viscous; 0.5 to 0.7% agar). The lowest swimming value has 3 (mm), while the highest value was 42.67 (mm) at addition of 10 % WCW in agar medium. The lowest swarming value has 7.66 (mm), while the highest value was 59.33 (mm) at addition of 10 % WCW in agar medium. In all experimental conditions, the addition of WCW to controls resulted an increase of up to 2.4 (swim) and 6.4 times (swarm) was observed. With the addition of WCW to the medium, an increase in both face and sliding motion was observed. However, these increases in the presence of NB medium are based on 60 mm. In the agar medium, according to the controls, when WCW was added, the sliding motion was 113 %, in the swimming motion 644 %; when WCW is added according to the controls in the NB medium, it is seen that there are 244 % increase in the swarming and 122 % increase in the swimming. The biggest difference is observed in the sliding motion (644 %) in the control (agar) (Figures 7-10). If the NB medium is compared with the controls, it is seen that there are 685% increase in swarming and 221 % increase in swimming (Figures 7-10).
Discussion
WCW contributed to the formation of cfu/ml, OD600 and biofilm, while causing a decrease in BFI values. In another published work we did, we used waste oil. When we compared the study with the whey trial, we reached higher values of cfu, OD600, biofilm and BFI with this study. The addition of WCW and increased ventilation (rinse) has resulted in increased value in all experimental conditions. Although the legacy BFI values also increased, this increase caused the increase to be below the control conditions. Therefore, the addition of WCW did not have a significant effect on BFI. The highest biofilm production occurred at 200 rpm, the highest number of cells; the cell density (live + dead) at bacterial density does not support this. It makes us think that the cells have been alive without division for a long time. Only 24 hours of time are used in our work. We interpreted the results by limiting it to this time slice. However, in the case of some similar works, over time zones of 72 hours were used [6,7,33].
For this reason, it was not possible to make a comparison. These studies have shown that whey wastes may be relatively better substrates for commercial production of biofilm and management of these wastes. There are also many studies supporting this [1- 3,6]. As a result, the addition of WCW causes a significant increase in both swarming and swimming. We did not find any study conducted by WCW in a similar way to the study we conducted in our article researches about the movement.
References
- Gomaa EZ (2013) Antimicrobial activity of a biosurfactant produced by Bacillus licheniformis strain M104 grown on Whey. Brazilian Archives of Biology and Technology 56(2): 259-268.
- Rodríguez-Pazo N, Vázquez-Araújo L, Pérez-Rodríguez N, Cortés-Diéguez S, Domínguez JM (2013) Cell-free supernatants obtained from fermentation of cheese whey hydrolyzates and phenylpyruvic acid by Lactobacillus plantarum as a source of antimicrobial compounds, bacteriocins, and natural aromas. Applied and Environmental Microbiology 171(4): 1042-1060.
- Garcia C, Rendueles M, Diaz M (2017) Synbiotic fermentation for the co-production of lactic and lactobionic acids from residual dairy whey. Biotechnology Progress 33(5): 1250-1256.
- Kishtaa OA, Guoc Y, Mofarrahic M, Stanac F, Landsa LC, Sabah NAH (2017) Pulmonary Pseudomonas aeruginosa infection induces autophagy and proteasome proteolytic pathways in skeletal muscles: effects of a pressurized whey protein-based diet in mice. Food Nutrition Research 61(1325309): 1-13.
- Wenzel J, Fuentes L, Cabezas A, Etchebehere C (2017) Microbial fuel cell coupled to biohydrogen reactor: a feasible technology to increase energy yield from cheese whey. Bioprocess and Biosystems Engineering 40(6): 807-819.
- De Giorgia S, Raddadia N, Fabbrib A, Gallina Toschib T, Fava F (2018) Potential use of ricotta cheese whey for the production of lactobionic acid by Pseudomonas taetrolens strains. New Biotechnology. 42: 71-76.
- Alonso S, Rendueles M, Díaz M (2013) Feeding strategies for enhanced lactobionic acid production from whey by Pseudomonas taetrolens. Bioresource Technology 134: 134-142.
- Overhage J, Lewenza S, Marr AK, Hancock REW (2007). Identification of genes ınvolved in swarming motility using a Pseudomonas aeruginosa PAO1 Mini-Tn5-lux Mutant Library. American Society for Microbiology 189(5): 2164-2169.
- Yeung ATY, Torfs ECW, Jamshidi F, Bains M, Wiegand I, Hancock REW, Overhage J (2009) Swarming of Pseudomonas aeruginosa is controlled by a broad spectrum of transcriptional regulators, ıncluding MetR Journal of Bacteriology 191(18): 5592-5602.
- O’May C, Tufenkji N (2011) The swarming motility of Pseudomonas aeruginosa ıs blocked by cranberry proanthocyanidins and other tannin-containing materials. Applied and Environmental Microbiology 77(9): 3061-3067.
- Caiazza NC, Merritt JH, Brothers KM, O’Toole GA (2007) Inverse regulation of biofilm formation and swarming motility by Pseudomonas aeruginosa PA14. Journal of Bacteriology 189(9): 3603-3612.
- Semmler ABT, Whitchurch CB, Mattick JS (1999) A re-examination of twitching motility in Pseudomonas aeruginosa. Microbiology 145: 2863-2873.
- Deziel E, Comeau Y, Villemur R (2001) Initiation of biofilm formation by Pseudomonas aeruginosa 57RP correlates with emergence of hyperpiliated and highly adherent phenotypic variants deficient in swimming, swarming, and twitching motilities. Journal of Bacteriology 183(4): 1195-1204.
- Tremblay J, Richardson AP, Lépine F, Déziel E (2007) Self-produced extracellular stimuli modulate the Pseudomonas aeruginosa swarming motility behaviour. Environmental Microbiology 9(10): 2622-2630.
- Overhage J, Bains M, Brazas MD, Hancock REW (2008) Swarming of Pseudomonas aeruginosa is a complex adaptation leading to ıncreased production of virulence factors and antibiotic resistance. Journal of Bacteriology 190(8): 2671-2679.
- Tremblay J, Déziel E (2008) Improving the reproducibility of Pseudomonas aeruginosa swarming motility assays. Journal of Basic Microbiology 48(6): 509-515.
- Copeland MF, Weibel DB (2009) Bacterial Swarming: A model system for studying dynamic selfassembly. NIH Public Access 5(6): 1174-1187.
- Morris JD, Hewitt JL, Wolfe LG, Kamatkar NG, Chapman SM, Diener JM, Courtney AJ, Matthew Leevy W, Shrout JD (2011) Imaging and analysis of Pseudomonas aeruginosa swarming and rhamnolipid production. Applied and Environmental Microbiology 77(23): 8310–8317.
- Wolska K, Szweda P, Lada K, Rytel E, Gucwa K, Kot B, Piechota M (2014) Motility activity, slime production, biofilm formation and genetic typing by ERIC-PCR for Pseudomonas aeruginosa strains isolated from bovine and other sources (human and environment). Polish Journal of Veterinary Sciences 17(2): 321-329.
- Murray TS, Kazmierczak BI (2006) FlhF is required for swimming and swarming in Pseudomonas aeruginosa. Journal of Bacteriology 188(19): 6995–7004.
- Alonso S, Rendueles M, Díaz M (2012) Role of dissolved oxygen availability on lactobionic acid production from whey by Pseudomonas taetrolens. Bioresource Technology 109: 140-147.
- Kim H, Park HD (2013) Ginger extract inhibits biofilm formation by Pseudomonas aeruginosa PA14. Plos One 8(9): 1-16.
- Wu C, Cheng Y, Yin H, Song X, Li W, et al. (2013) Oxygen promotes biofilm formation of Shewanella putrefaciens CN32 through a diguanylate cyclase and an adhesin. Nature Science Reports 3(1945): 1-7.
- Dos Santos Goncalves M, Delattre C, Balestrino D, Charbonne N, Elboutachfaiti R, et al. (2014) Anti-biofilm activity: a function of Klebsiella pneumoniae capsular polysaccharide. Plos One 9(6): 1-12.
- Adamus-Bialek W, Kubiak A, Czerwonka G (2015) Analysis of uropathogenic Escherichia coli biofilm formation under different growth conditions. ACTA Biochimica Polonica 62(4): 765-771.
- Crespo A, Pedraz L, Astola J, Torrents E (2016) Pseudomonas aeruginosa exhibits deficient biofilm formation in the absence of class II and III ribonucleotide reductases due to hindered anaerobic growth. Frontiers in Microbiology 7(688): 1-14.
- Castro J, Alves P, Sousa C, Cereija T, França  , et al. (2015) Using an in-vitro biofilm model to assess the virulence potential of bacterial vaginosis or non- bacterial vaginosis Gardnerella vaginalis isolates. Scientific Reports 5(11640): 1-10.
- Naves P, del Prado GL, Huelves MG, Ruiz V, Blanco J, et al. (2008) Measurement of biofilm formation by clinical isolates of Escherichia coli is method-dependent. Journal of Applied Microbiology 105: 585-590.
- Berlanga M, Gomez-Perez L, Guerrero R (2017) Biofilm formation and antibiotic susceptibility in dispersed cells versus planktonic cells from clinical, industry and environmental origins. Antonie van Leeuwenhoek 110(12): 1691-1704.
- Inoue T, Shingaki R, Fukui K (2008) Inhibition of swarming motility of Pseudomonas aeruginosa by branched-chain fatty acids. FEMS Microbiology Letters 281: 81-86.
- Murray TS, Kazmierczak BI (2008) Pseudomonas aeruginosa exhibits sliding motility in the absence of type IV pili and flagella. Journal of Bacteriology 190 (8): 2700-2708.
- Fuente-Núñez C, Korolik V, Bains M, Nguyen U, Breidenstein EBM, et al. (2012) Inhibition of bacterial biofilm formation and swarming motility by a small synthetic cationic peptide. Antimicrobial Agents and Chemotherapy 56(5): 2696-2704.
- Lo R, Xue T, Weeks M, Turner MS, Bansal N (2016) Inhibition of bacterial growth in sweet cheese whey by carbon dioxide as determined by culture-independent community profiling. International Journal of Food Microbiology 217: 20-28.
 Research Article
Research Article