Keywords: Biomarker; Metastatic colorectal cancer; Predictive
Introduction
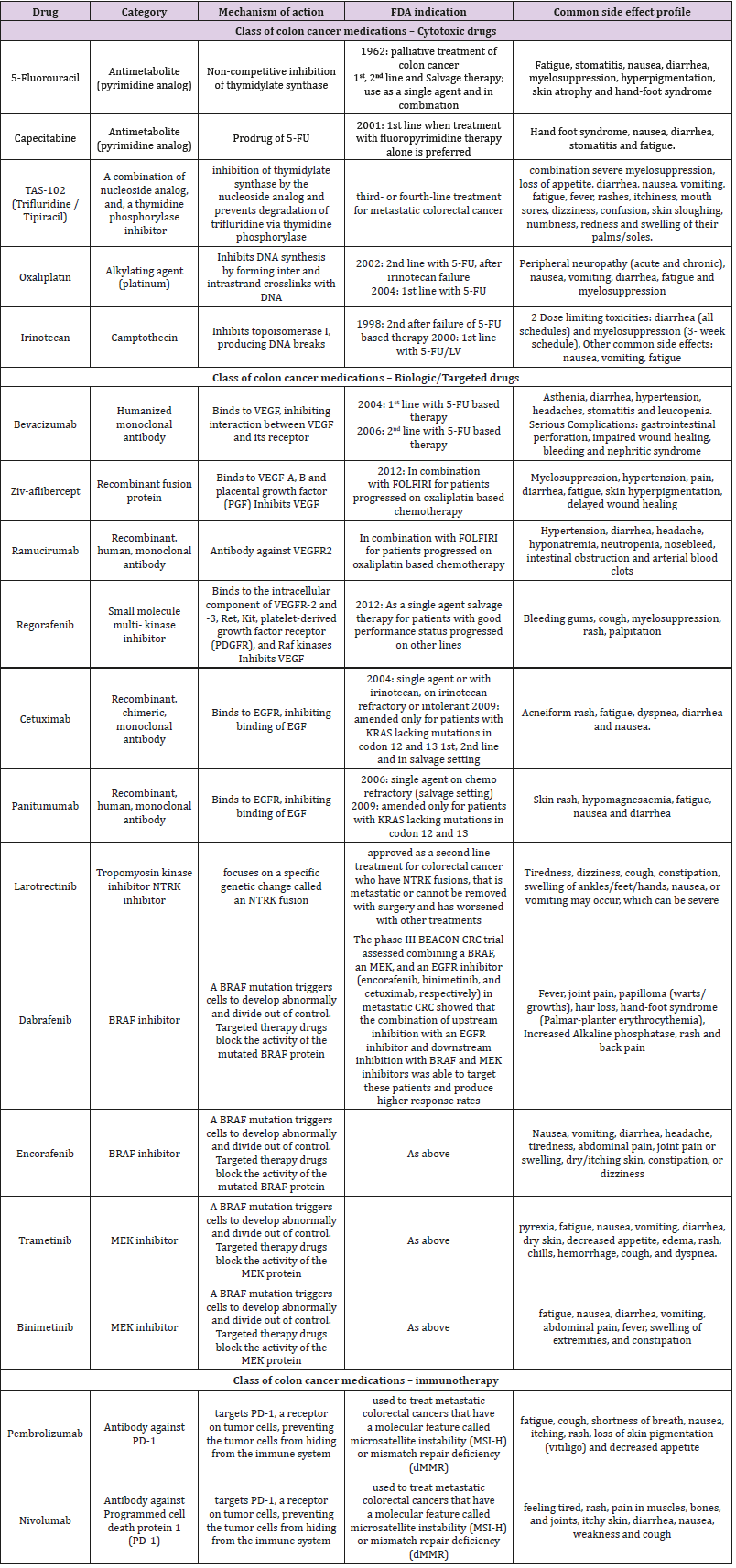
Colorectal cancer is the third most common cancer detected in both men and women in the United States. As per American Cancer Society, 101,420 new cases of colon and 44,180 new cases of rectal cancer are expected in the United States, for the year 2019. The lifetime risk of developing CRC is expected to be 1 out of 22 (4.49%) for men and 1 out of 24 (4.15%) for women. In terms of mortality, CRC is the second leading cause of cancer related mortality in both men and women, accounting for 51,020 deaths in 2019 [1,2]. Historically, 5-fluorouracil (5-FU) has been the standard of care for metastatic CRC (mCRC) but following the approval of cytotoxic agents and several targeted therapies in the last two decades, the landscape of treatment of mCRC has changed, leading to increased tumor response and patient survival rates [3]. Cytotoxic agents have been combined with biologic agents in the current standard of care for mCRC. A total of ten different classes of drugs (three classes of cytotoxic agents, six classes of biologic agents, namely anti EGFR, anti-VEGF, multitargeted tyrosine kinase inhibitors, NTRK inhibitors, BRAF inhibitors, MEK inhibitors and one class of immunotherapy) are currently being used for the treatment of mCRC. An overview of all the different drugs that have been approved by the US Food and Drug Administration (USFDA) for use in the management of mCRC, with their mechanism of action and adverse effect profile has been described in Table 1 [4-6].
Predictive Biomarkers
Due to the heterogeneity in mCRC, not all the patients may benefit from the same treatment strategy. Predictive biomarkers can help us to know what will be the likely response of a patient to a particular therapy. Predictive biomarkers can guide the selection of the best treatment strategy for individual patients.
Cytotoxic Agents
We have seen minimal improvements in the biomarkers driven cytotoxic strategies in the past decade. Most known predictive biomarkers for cytotoxic agents are as follows.
DPD Deficiency: Dihydropyrimidine dehydrogenase (DPYD) is a gene encode for the enzyme, dihydropyrimidine dehydrogenase (DPD), a rate-limiting enzyme in the fluoropyrimidine metabolism pathway. Polymorphism in the DPYD gene leads to varying levels of tolerating the fluoropyrimidines (fluorouracil and its oral formulation capecitabine); while a partial or complete deficiency in the DPD activity puts the patient at high risk for developing lifethreatening adverse events [7-15]. Particularly the alleles DPYD*2A and A2846T are associated with developing life-threatening severe toxicities in patients receiving fluorouracil or capecitabine therapy [16,17].
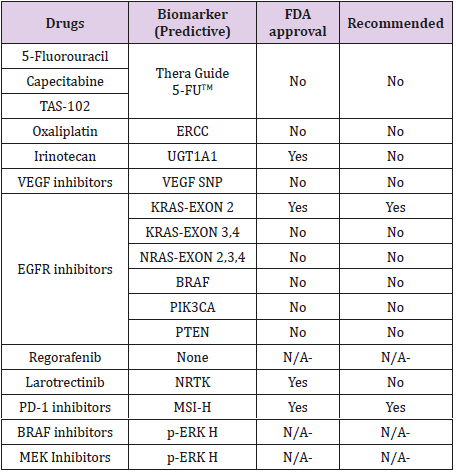
TYMS Gene Variations: 5-FU potentially inhibits the enzyme thymidylate synthetase (TS), encoded by the thymidylate synthetase gene (TYMS). TYMS gene has polymorphism for double (2R) or triple (3R) repeat in its promotor region of the gene, which affects the TS levels. It has been observed that patients who are homozygous 2R/2R have low TS levels and may be at higher risk for toxicities up to 1.4 to 2.4 times with 5-FU [18-19]. Currently, there is a commercially available test, namely, “Thera Guide 5-FU”. This test may be used to analyze both DYPD gene and TYMS gene polymorphisms but is not routinely seen being done in clinical practice as it is expensive and takes around 7days to report the results.
Micro RNA-143 (miR-143): MicroRNAs regulate the expression levels of other genes by multiple mechanisms. It is observed that low miR-143 expression was associated with longer progression-free survival (PFS) in patients on capecitabine therapy [20]. Contrastingly high miR-143 expression showed longer PFS in patients with 5-FU [21]. It could be understood that the function of miR-143 differs under different circumstances like 5-FU vs. capecitabine. However, its credibility as a biomarker can be established only after further studies. Hence, it is a potential predictive biomarker.
UGT1A1 Polymorphisms: The active metabolite of irinotecan is metabolized by enzyme uridine diphospho-glucuronosyltransferase 1A1 (UGT1A1). This enzyme shows polymorphism with reduced enzymatic activity among those who inherit the UGT1A1*28 allele. A study demonstrated that patients with homozygous and to some extent, heterozygous for allele UGT1A1*28 developed more severe toxicities with irinotecan therapy. However, it also showed that these patients have better response rates and PFS compared with another allele UGT1A*1 [22].
ABCC5 and ABCG1 Polymorphisms – Irinotecan Toxicity: Polymorphism in the ATP-binding cassette (ABC) genes that participate in the pharmacokinetics of irinotecan were studied. It was observed that the polymorphic transporter genes ABCG1 and ABCC5 were associated with increased risk of developing neutropenia and diarrhea, respectively, in patients treated with irinotecan. This biomarker can potentially further personalize the FOLFIRI treatment of mCRC patients [23].
Topoisomerase 1 Copy Number Alterations: It is known that FOLFIRI and FOLFOX are considered as equally valid as first-line therapy in the mCRC. However, clinicians choose one from their experience or one that they are comfortable managing their toxicities. It has been observed in a study that there is a significant association between the increasing copy number (CN) of topoisomerase 1 (TOP1) and response to irinotecan monotherapy [24] thus suggesting FOLFIRI over FOLFOX in these patients.
Biologic or Targeted Agents
Mutations in RAS
Epidermal Growth Factor (EGF) binding the EGF receptor (EGFR) plays a pivotal role in the regulation of cell differentiation and proliferation by activation of RAS-RAF-MEK-ERK-MAP kinase pathway. Many studies have established that the presence of RAS (KRAS) mutations is predictive of lack of clinical benefit to monoclonal antibodies that bind to EGFR. Besides, they have proven to be harmful when combined with oxaliplatin [25,26]. Therefore, it is recommended that panitumumab and cetuximab (monoclonal antibodies against EGFR) are utilized only in patients with wild type RAS (wt RAS). Results from the PRIME study demonstrated that not only the exon 2 (codon 12 and 13) with seven specific mutations that account for over 90% of KRAS mutations but also KRAS exon 3 (codon 61) and exon 4 (codon 117 and 146), and NRAS mutations also predict for nonresponse to EGFR inhibitor therapy [27]. In addition, it was observed that some patients with wt RAS also did not benefit from anti-EGFR therapy. On further studies, patients with wt KRAS, some miRNAs predicted response to anti-EGFR treatment. Low expression of miR-1881a has been associated with unfavorable outcomes [28-29]. Two tests are FDA approved to test for the mutations in RAS, which include the Cobas KRAS mutation test and Praxis Extended RAS Panel.
BRAF Mutation: The RAF is one of the downstream mediators of the EGFR signaling pathway. Two meta-analyses have revealed that the patients treated with EGFR inhibitors did not show any improvement in terms of PFS or overall survival when compared to chemotherapy alone [30-31]. Based on these results, EGFR inhibitors are not recommended in BRAF mutant patients.
PTPRT and PTPRD: It is observed in a study that deleterious mutation in receptor-type tyrosine-protein phosphatase T (PTPRT), a phosphatase involved in JAK/STAT signaling pathway and its related gene PTPRD correlated with the resistance towards bevacizumab therapy [32]. However, the study involved a tiny sample; hence, more extensive studies must be done to establish the observed effect.
NTRK: In 2018, Larotrectinib, a kinase inhibitor, first of its kind approved for the treatment of malignancies with neurotrophic tyrosine kinase receptor (NTRK) gene fusion without a known acquired resistance mutation, irrespective of the tissue origin of the tumor. NTRK gene fusion happens in only one percent of patients with mCRC; however, in these patients, larotrectinib is a valuable option. Its rarity in mCRC makes it expensive to test and difficult to detect [33].
Immunotherapy Agents
dMMR/MSI-H: Deficient mismatch repair/ Microsatellite instability-high are indicators of genomic instability. Recently, the National Comprehensive Cancer Network (NCCN) guidelines have included nivolumab or pembrolizumab for patients who have progressed following treatment with cytotoxic agents or who are not eligible for cytotoxic combinations. These agents may be used as a second or third line for patients with dMMR/MSI-H positive mCRC [33].
Prognostic Markers
Prognostic markers are biomarkers that give overall progress of a disease in a patient population. Previously, the classical prognostic markers in oncology include the size of the tumor, stage, and presence of metastasis. With the advent of modern molecular techniques and genomic sequencing technology, molecular biomarkers are being discovered which can establish some prognostic value. Table 2 lists out some of the prognostic markers that have been studied and can potentially be used in the clinical practice.
Conclusion
Although many potential predictive and prognostic biomarkers are being reported in various studies, very few of them are used in the clinical practice due to varying reasons. More clinical studies must be done to identify newer potential biomarkers and to establish the clinical validity of some know biomarkers so that they can be used in clinical practice to individualized treatment options and predict the outcomes.
References
- (2019) American Cancer Society. Cancer Facts & Figures.
- Siegel RL, Miller KD, Jemal A (2019) Cancer statistics, 2019. CA Cancer J Clin69(1): 7-34.
- Patil H, Saxena SG, Barrow CJ, Kanwar JR, Kapat A, et al. (2017) Chasing the personalized medicine dream through biomarker validation in colorectal cancer. Drug Discov Today 22(1): 111-119.
- Aparo S, Goel S (2012) Evolvement of the treatment paradigm for metastatic colon cancer. From chemotherapy to targeted therapy. Crit Rev Oncol Hematol 83(1): 47-58.
- Nguyen MT, Weinberg DS (2016) Biomarkers in Colorectal Cancer Screening. J Natl Compr Cancer Netw 14(8): 1033-1040.
- (2017) US. Food and Drug Administration.
- Diasio RB, Beavers TL, Carpenter JT (1988) Familial deficiency of dihydropyrimidine dehydrogenase. Biochemical basis for familial pyrimidinemia and severe 5-fluorouracil-induced toxicity. J Clin Invest 81(1): 47-51.
- Maring JG, van Kuilenburg ABP, Haasjes J, H Piersma, HJ M Groen, et al. (2002) Reduced 5-FU clearance in a patient with low DPD activity due to heterozygosity for a mutant allele of the DPYD gene. Br J Cancer 86(7): 1028-1033.
- Ezzeldin H, Johnson MR, Okamoto Y, Diasio R (2003) Denaturing high performance liquid chromatography analysis of the DPYD gene in patients with lethal 5-fluorouracil toxicity. Clin Cancer Res 9(8): 3021-3028.
- Harris BE, Carpenter JT, Diasio RB (1991) Severe 5-fluorouracil toxicity secondary to dihydropyrimidine dehydrogenase deficiency. A potentially more common pharmacogenetic syndrome. Cancer 68(3): 499-501.
- Takimoto CH, Lu ZH, Zhang R, Liang MD, Larson LV, et al. (1996) Severe neurotoxicity following 5-fluorouracil-based chemotherapy in a patient with dihydropyrimidine dehydrogenase deficiency. Clin Cancer Res 2(3): 477-481.
- Ezzeldin H, Diasio R (2004) Dihydropyrimidine Dehydrogenase Deficiency, a Pharmacogenetic Syndrome Associated with Potentially Life-Threatening Toxicity Following 5-Fluorouracil Administration. Clin Colorectal Cancer 4(3): 181-189.
- Diasio RB, Johnson MR (1999) Dihydropyrimidine dehydrogenase: its role in 5-fluorouracil clinical toxicity and tumor resistance. Clin Cancer Res 5(10): 2672-2673.
- Van Kuilenburg ABP, Meinsma R, Zonnenberg BA, Baas F, Matsuda K, et al. (2003) Dihydropyrimidinase deficiency and severe 5-fluorouracil toxicity. Clin Cancer Res 9(12): 4363-4367.
- Yang CG, Ciccolini J, Blesius A, Laetitia Dahan, Danielle Bagarry-Liegey, et al. (2011) DPD-based adaptive dosing of 5-FU in patients with head and neck cancer: impact on treatment efficacy and toxicity. Cancer Chemother Pharmacol 67(1): 49-56.
- Deenen MJ, Meulendijks D, Cats A, Marjolein K Sechterberger, Johan L Severens, et al. (2016) Upfront Genotyping of DPYD*2A to Individualize Fluoropyrimidine Therapy: A Safety and Cost Analysis. J Clin Oncol 34(3): 227-234.
- Henricks LM, Lunenburg CATC, de Man FM, Meulendijks D, Frederix GWJ, et al. (2018) DPYD genotype-guided dose individualisation of fluoropyrimidine therapy in patients with cancer: a prospective safety analysis. Lancet Oncol 19(11): 1459-1467.
- Ichikawa W, Takehiro Takahashi, Kenichi Suto, Yasutsuna Sasaki, Renzo Hirayama (2006) Orotate Phosphoribosyltransferase Gene Polymorphism Predicts Toxicity in Patients Treated with Bolus 5-Fluorouracil Regimen. Clin Cancer Res 12(13): 3928-3934.
- Lecomte T, JeanMarc Ferraz, Franck Zinzindohoué, Marie Anne Loriot, David Alexandre Tregouet, et al. (2004) Thymidylate Synthase Gene Polymorphism Predicts Toxicity in Colorectal Cancer Patients Receiving 5-Fluorouracil-based Chemotherapy. Clin Cancer Res 10(17): 5880-5888.
- Simmer F, Venderbosch S, Dijkstra JR, Elisa M. Vink-Börger, Claudius Faber, et al. (2015) MicroRNA-143 is a putative predictive factor for the response to fluoropyrimidine-based chemotherapy in patients with metastatic colorectal cancer. Oncotarget 6(26): 22996-23007.
- Borralho PM, Kren BT, Castro RE, Moreira da Silva IB, Steer CJ, et al. MicroRNA-143 reduces viability and increases sensitivity to 5-fluorouracil in HCT116 human colorectal cancer cells. FEBS J 276(22): 6689-6700.
- Toffoli G, Cecchin E, Corona G, Antonio Russo , Angela Buonadonna, et al. (2006) The Role of UGT1A1*28 Polymorphism in the Pharmacodynamics and Pharmacokinetics of Irinotecan in Patients With Metastatic Colorectal Cancer. J Clin Oncol 24(19): 3061-3068.
- Chen S, Villeneuve L, Jonker D, Couture F, Laverdière I, et al. (2015) ABCC5 and ABCG1 polymorphisms predict irinotecan-induced severe toxicity in metastatic colorectal cancer patients. Pharmacogenet Genomics 25(12): 573-583.
- Palshof JA, Høgdall EVS, Poulsen TS, Dorte Linnemann, Benny Vittrup Jensen, et al. (2017) Topoisomerase I copy number alterations as biomarker for irinotecan efficacy in metastatic colorectal cancer. BMC Cancer 17(1): 48.
- Han CB, Li F, Ma JT, Zou HW (2012) Concordant KRAS Mutations in Primary and Metastatic Colorectal Cancer Tissue Specimens: A Meta-Analysis and Systematic Review. Cancer Invest 30(10): 741-747.
- Peeters M, Kafatos G, Taylor A, Gastanaga VM, Oliner KS, et al. (2015) Prevalence of RAS mutations and individual variation patterns among patients with metastatic colorectal cancer: A pooled analysis of randomised controlled trials. Eur J Cancer 51(13): 1704-1713.
- Douillard JY, Siena S, Cassidy J, J Tabernero, R Burkes, et al. (2014) Final results from PRIME: randomized phase III study of panitumumab with FOLFOX4 for first-line treatment of metastatic colorectal cancer. Ann Oncol 25(7): 1346-1355.
- Cappuzzo F, Sacconi A, Landi L, Ludovini V, Biagioni F, et al. (2014) MicroRNA Signature in Metastatic Colorectal Cancer Patients Treated with Anti-EGFR Monoclonal Antibodies. Clin Colorectal Cancer 13(1): 37-45.e4.
- Pichler M, Winter E, Ress AL, Thomas Bauernhofer, Armin Gerger, et al. (2014) miR-181a is associated with poor clinical outcome in patients with colorectal cancer treated with EGFR inhibitor. J Clin Pathol 67(3): 198-203.
- Pietrantonio F, Petrelli F, Coinu A, Di Bartolomeo M, Borgonovo K, et al. (2015) Predictive role of BRAF mutations in patients with advanced colorectal cancer receiving cetuximab and panitumumab: A meta-analysis. Eur J Cancer 51(5): 587-594.
- Therkildsen C, Bergmann TK, Henrichsen Schnack T, Ladelund S, Nilbert M (2014) The predictive value of KRAS, NRAS, BRAF, PIK3CA and PTEN for anti-EGFR treatment in metastatic colorectal cancer: A systematic review and meta-analysis. Acta Oncol 53(7): 852-864.
- Hsu HC, Lapke N, Chen SJ, Lu YJ, Jhou RS, et al. (2018) PTPRT and PTPRD Deleterious Mutations and Deletion Predict Bevacizumab Resistance in Metastatic Colorectal Cancer Patients. Cancers (Basel) 10(9): 314.
- Messersmith WA (2019) NCCN Guidelines Updates: Management of Metastatic Colorectal Cancer. J Natl Compr Canc Netw 17(5.5): 599-601.
 Mini Review
Mini Review