ABSTRACT
The rapidly spreading Severe Acute Respiratory Syndrome Coronavirus 2 (SARSCoV- 2) has inflicted numerous patients and deaths in human inhabiting city-states worldwide. Due to its zoonotic nature, not only humans are susceptible to the recent threat. Non-human wildlife such as white-tailed deer (Odocoileus virginianus), the Père David’s deer (Elaphurus davidianus), and the typical roe deer (Capreolus capreolus) are all highly susceptible to SARS-CoV-2 due to the virus targeting highly conserved genes such as host angiotensin-converting enzyme 2 (ACE-2) receptors. While the virus was vastly spreading globally within human patients, wildlife was also contacted via viral residue discarded in local cities or direct contact with infected patients. The current situation of cervids and capreolus species are reported to be inflicted by SARS-CoV-2 vastly throughout multiple countries. The ability to contain the coronavirus and act as both a reservoir and a method to successfully transmission the virus vertically to future generations will indefinitely enable SARS-CoV-2 to develop novel mutations and in turn unleash another possible zoonotic outbreak.
Keywords: SARS-CoV-2; Wildlife; Human; Cervid; Zoonosis
Introduction
Severe Acute Respiratory Syndrome Coronavirus 2 (SARSCoV- 2) is a positive sense single-stranded RNA virus that constantly threatens the public health [1]. After the initial identification, the pandemic had spread to regions far across its originated continent claiming more than 5.9 million deaths and infecting more than 433 million people around the world [2]. The infection alone did not lead the virus as a global threat. The ability to transform into multiple subtype variants due to its RNA nature made governments and health care departments more challenging to successfully identify and cure infected patients. While the current state of SARS-CoV-2 is a major concern, the event of its emergence is also important because the transmission itself was zoonotic. SARSCoV- 2 is categorized within the severe acute respiratory syndrome related Coronavirus (SARSr-CoV) of the family Coronaviridae [3]. In attempts to discover which animal started the infection, reports indicated a genome sequence of one of the SARr-CoV which resembled 96% of the whole RaTG13 gene within SARS-CoV-2 [4]. The report suggested hard evidence of viral genome sequences from bats were the origin of the initial infection. Since the report was released, several other bat viral genome sequences related to SARS-CoV-2 have been sighted from different regions in China and Japan [5-8]. Considering previous reports, it may have been inevitable of cross-infection of the SARS-CoV-2 virus from nonhuman mammals. As the infection continue among the human population, other mammals were also in the risk of exposure by any contacts with human patients. SARS-CoV-2 targets the host angiotensin-converting enzyme 2 (ACE-2) receptor to tether cells [9]. Interestingly, ACE-2 receptor are highly conserved genes which are observed in various animal species [10]. As a result, various incidents regarding dogs, cats, zoo animals, and even farmed minks were documented as a SARS-CoV-2 infection [11-14]. While domestic and industrial mammals were the first to be infected due to numerous human contacts, the continuous widespread has led to further spillovers regarding local wildlife [15]. Among the indigenous wildlife, capreolus and cervid species were reported to be highly susceptible to SARS-CoV-2, especially white-tailed deer (Odocoileus virginianus), the Père David’s deer (Elaphurus davidianus), and the typical roe deer (Capreolus capreolus) [16]. This review is to aim on the current SARS-CoV-2 situation in wild capreolus and cervid species and the potential zoonotic backlash it will bring to human public health care.
Methods of Transmission from Human to Wildlife
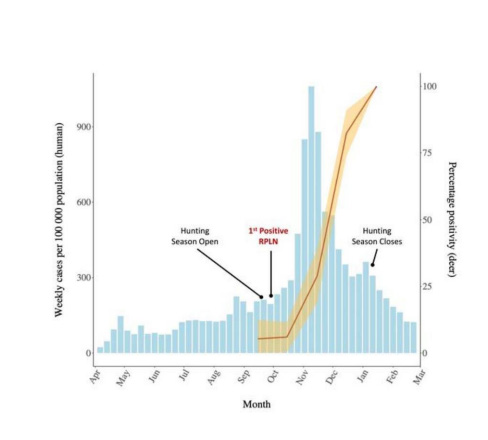
When compared to local domestic and industrial animals, wildlife located far from suburbs are considered less hazardous to confront direct SARS-CoV-2 infection. However, reports suggest a high possibility that infected patients shed SARS-CoV-2 within feces [17]. Recent research on other Coronavirus subtypes such as Middle East Respiratory Syndrome (MERS) report detection rates of 14.6% and 2.4% within fecal and urine samples [18]. The predecessor SARS-CoV inflicted patients experienced diarrhea and SARS-CoV RNA was detected within the stool samples and small intestines [19]. The SARS-CoV virus was able to remain contact and fully functional in diarrheal samples for a maximum 4 days at room temperature [20]. Similar to its former subtypes, a recent report discovered a U. S. patient infected with SARS-CoV-2 shed SARS-CoV-2 RNA in their feces [21]. More than 50% 96 patients’ feces samples collected over 31 days detected SARS-CoV-2 [22]. The research proves that a portion of the infected patients are able to shed SARS-CoV-2 virus via diarrheal feces. The disposed virus derived from populated suburbs, hospitals, and airports will be transported to various sewage systems. Mistreated water has the potential to carry SARS-CoV-2 RNA and act as a potential vehicle to expose the virus directly to the natural aquatic environment [23- 25]. Coronavirus has been known to lose its initial infectivity when exposed in water due to its lipid envelope structure [26]. However, depending on the viral subtype and condition of the water such as temperature, Human derived Coronavirus are known to survive more than 500 days [27]. This provides ample time for wandering roe deer and other cervids to be contacted by the virus and be infected via the gastrointestinal organs. The other major method of a spillover would be due to seasonal hunting. Studies regarding retropharyngeal lymph node samples of wild deer discovered a sharp increase in positive SARS-CoV-2 RNA signatures when the hunting season started on September 19, 2020, until January 10, 2021 [28]. The study in Iowa suggests major influx of people during the hunting season left various SARS-CoV-2 containing residue infecting the Southeastern area (Figure 1) Suresh V. Kuchipudi, et al. [28]. The virus contacted deer acted as a viral reservoir to actively spread to other regions, gradually infecting in all directions. This state limited situation later developed to regional crossing inflicting SARS-CoV-2 infection in other states.
Figure 1: Epidemic curve showing SARS-CoV-2 weekly cases (per 100,000) in humans and the monthly change in SARS-CoV-2 positivity in White-tailed deer in Iowa. Adapted from Multiple spillovers and onward transmission of SARS-Cov-2 in free-living and captive White-tailed deer (Odocoileus virginianus) by Suresh V. Kuchipudi, 2022, Proc Natl Acad Sci 119(6).
Current State of Capreolus and Cervid Species Infection
While SARS-CoV-2 was widespread in major human populated regions, SARS-CoV-2 wildlife dissemination were not yet sighted. The trend of the viral infection tends to have a gap of one year in the viral progression. As of today, situation of SARS-CoV-2 infecting various cervid and capreolus species is spotted in almost every continent. Vast samples of deer originating from different countries in North America such as Ohio and Maryland in the United States and Canada have detected SARS-CoV-2 RNA [29]. SARS-CoV-2 detection efforts are also increased in Eurasian countries such as northern Fennoscandia, Iceland, and Eastern Russia [30]. Compared to the western regions, reports regarding cervid and capreolus species inhabiting eastern countries are relatively short. Main reports are focused on the direct culprit which started the viral outbreak. Japan, China, and Singapore reported positive identification of the same SARS-CoV-2 strand containing bats. This strongly predicts there will be high chance of local deer or other capreolus species exposed to SARS-CoV-2 residue and naturally be infected.
Viral Characteristics and Variants among the Infected Cervid Species
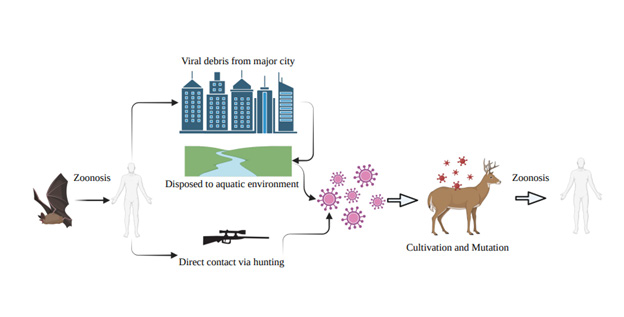
SARS-CoV-2 drastically developed within the human patients by transforming through a series of mutations later discovered as novel variants of the original virus. Within 2019 to 2022, major SNP mutations named as Alpha, Beta, Gamma, Delta, and the latest Omicron variant quickly spread to heavily populated human settlements. Similarly, local wildlife has also contacted the gradually mutating virus. It is crucial for both the infected deer and human patient to be identified which subtype of SARS-CoV-2 as every variant mostly contains variant specific point mutations. For example, the previous alpha variant was identified as a novel variant due to mutations such as N501Y, D614G, and P681H. This enabled the mutated spike proteins of SARS-CoV-2 to improve its ability to bind cellular receptors of host cells [31]. The more recent variant Delta and Omicron contained more unique point mutations such as the T478K, P681R, and K417N mutation in Delta and E484D, P812R, and Q954H in Omicron variants [32,33]. While the alpha and other previously found variants are hindering within the human population, wild deer and other capreolus species are in the relatively early stages of dissemination. What is more concerning with the infected deer is that not only does the animals act as a suitable reservoir for SARS-CoV-2, but also the ability to transmission the virus from doe to fetus. Recent reports suggest adult white-tailed deer can transmit the Alpha variant of SARSCoV- 2 vertically [34]. The virus’s ability of vertical transmission in utero have been documented in human pregnant patients [35], but the ability of other wildlife mammals to pass on previous variants will indefinitely influence future SARS-CoV-2 development. Infected cervids will contain the virus through generations and may eventually harbor novel mutations that will in turn trigger yet another zoonosis to unsuspecting humans (Figure 2).
Figure 2: Schematics of SARS-CoV-2 zoonosis and pathways of cervid infection leading to potential reverse zoonosis.
Conclusion
Studies regarding the current spread of SARS-CoV-2 infection in wild cervid or capreolus species are yet to be finished. Further sequencing and observation will be required to successfully document how far wildlife have been exposed to the zoonotic virus that may prevent another, perhaps more serious outbreak of reverse zoonosis.
Declarations
Ethical Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Data Availability Statement
The datasets used and/or analyzed during the current study available from the corresponding author on reasonable request.
Competing Interests
The author declares no competing interests.
Funding
This work was supported by the Precision Medical Device Hyperconvergence project funded by the Ministry of Trade, Industry and Energy (#NP2020-0120).
Authors’ Contributions
Hyoung-Min Park contributed in review design, manuscript writing, and overall process.
Acknowledgement
Not applicable.
References
- Wang C, Peter W Horby, Frederick G Hayden, George F Gao (2020) A novel coronavirus outbreak of global health concern. Lancet 395(10223): 470-473.
- (2022) WHO. Weekly epidemiological update on COVID-19.
- (2020) Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol 5(4): 536-544.
- Zhou P, Xing-Lou Yang, Xian Guang Wang, Ben Hu, Lei Zhang, et al. (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579(7798): 270-273.
- Zhou H, Xing Chen, Tao Hu, Juan Li, Hao Song, et al. (2020) A novel bat coronavirus closely related to SARS-CoV-2 contains natural insertions at the S1/S2 cleavage site of the spike protein. Current biology 30(11): 2196-2203. e3.
- Murakami S, Tomoya Kitamura, Jin Suzuki, Ryouta Sato, Toshiki Aoi, et al. (2020) Detection and Characterization of Bat Sarbecovirus Phylogenetically Related to SARS-CoV-2, Japan. Emerg Infect Dis 26(12): 3025-3029.
- Xiao K, Junqiong Zhai, Yaoyu Feng, Niu Zhou, Xu Zhang, et al. (2020) Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins. Nature 583(7815): 286-289.
- Lam TT, Na Jia, Ya Wei Zhang, Marcus Ho Hin Shum, Jia-Fu Jiang, et al. (2020) Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature 583(7815): 282-285.
- Wan Y, Jian Shang, Rachel Graham, Ralph S. Baric, Fang Li (2020) Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J Virol 94(7).
- Damas J, Graham M Hughes, Kathleen C Keough, Corrie A Painter, Nicole S Persky, et al. (2020) Broad Host Range of SARS-CoV-2 Predicted by Comparative and Structural Analysis of ACE2 in Vertebrates. bioRxiv 117(36): 22311-22322.
- Oreshkova N, Robert Jan Molenaar, Sandra Vreman, Frank Harders, Bas B Oude Munnink, et al. (2020) SARS-CoV-2 infection in farmed minks, the Netherlands, April and May 2020. Euro Surveill 25(23): 2001005.
- Fenollar F, Oleg Mediannikov, Max Maurin, Christian Devaux, Philippe Colson, et al. (2021) Mink, SARS-CoV-2, and the Human-Animal Interface. Front Microbiol 12: 663815.
- Buonavoglia C, Nicola Decaro, Vito Martella, Gabriella Elia, Marco Campolo, et al. (2006) Canine coronavirus highly pathogenic for dogs. Emerg Infect Dis 12(3): 492-494.
- Sharun K, Kuldeep Dhama, Abhijit M Pawde, Christian Gortázar, Ruchi Tiwari, et al. (2021) SARS-CoV-2 in animals: potential for unknown reservoir hosts and public health implications. Vet Q 41(1): 181-201.
- Weiss SR, S Navas Martin (2005) Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol Mol Biol Rev 69(4): 635-664.
- Palmer MV, Mathias Martins, Shollie Falkenberg, Alexandra Buckley, Leonardo C Caserta, et al. (2021) Susceptibility of white-tailed deer (Odocoileus virginianus) to SARS-CoV-2. J Virol 95(11): e00083-21.
- Franklin AB, SN Bevins (2020) Spillover of SARS-CoV-2 into novel wild hosts in North America: A conceptual model for perpetuation of the pathogen. Sci Total Environ 733: 139358.
- Corman VM, Ali M Albarrak, Ali Senosi Omrani, Mohammed M Albarrak, Mohamed Elamin Farah, et al. (2016) Viral Shedding and Antibody Response in 37 Patients with Middle East Respiratory Syndrome Coronavirus Infection. Clin Infect Dis 62(4): 477-483.
- Leung WK, Ka-Fai To, Paul K S Chan, Henry L Y Chan, Alan K L Wu, et al. (2003) Enteric involvement of severe acute respiratory syndrome-associated coronavirus infection. Gastroenterology 125(4): 1011-1017.
- Lai MY, PK Cheng, WW Lim (2005) Survival of severe acute respiratory syndrome coronavirus. Clin Infect Dis 41(7): e67-71.
- Holshue ML, Chas DeBolt, Scott Lindquist, Kathy H Lofy, John Wiesman, et al. (2020) First Case of 2019 Novel Coronavirus in the United States. N Engl J Med 382(10): 929-936.
- Zheng S, Jian Fan, Fei Yu, Baihuan Feng, Bin Lou, et al. (2020) Viral load dynamics and disease severity in patients infected with SARS-CoV-2 in Zhejiang province, China, January-March 2020: retrospective cohort study. BMJ 369: m1443.
- Ahmed W, Nicola Angel, Janette Edson, Kyle Bibby, Aaron Bivins, et al. (2020) First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community. Sci Total Environ 728: 138764.
- Medema G, Heijnen Leo, Elsinga Goffe, Italiaander Ronald, Brouwer Anke (2020) Presence of SARS-Coronavirus-2 RNA in sewage and correlation with reported COVID-19 prevalence in the early stage of the epidemic in the Netherlands. Environmental Science & Technology Letters 7(7): 511-516.
- Wu F, Amy Xiao, Jianbo Zhang, Katya Moniz, Noriko Endo, et al. (2020) SARS-CoV-2 titers in wastewater foreshadow dynamics and clinical presentation of new COVID-19 cases. medRxiv.
- Schoeman D, BC Fielding (2019) Coronavirus envelope protein: current knowledge. Virol J 16(1): 69.
- Gundy PM, CP Gerba, IL Pepper (2009) Survival of coronaviruses in water and wastewater. Food and Environmental Virology 1(1): 10-14.
- Kuchipudi SV, Meera Surendran Nair, Rachel M Ruden, Vivek Kapur, Katriina Willgert, et al. (2022) Multiple spillovers from humans and onward transmission of SARS-CoV-2 in white-tailed deer. Proc Natl Acad Sci U S A 119(6): e2121644119.
- Hale VL, Patricia M Dennis, Dillon S McBride, Devra Huey, Margot Ehrlich, et al. (2022) SARS-CoV-2 infection in free-ranging white-tailed deer. Nature 602(7897): 481-486.
- Sanchez Romano J, Anna Omazic, Mikael Leijon, Åsa Hagström, Morten Tryland, et al. (2021) Screening of Eurasian Tundra Reindeer for Viral Sequences by Next-Generation Sequencing. Int J Environ Res Public Health 18(12): 6561.
- Peiffer Smadja N, Antoine Bridier Nahmias, Valentine Marie Ferré, Mathilde Garé, Christophe Rioux, et al. (2021) Emergence of E484K Mutation Following Bamlanivimab Monotherapy among High-Risk Patients Infected with the Alpha Variant of SARS-CoV-2. Viruses 13(8): 1642.
- Cherian S, Varsha Potdar, Santosh Jadhav, Pragya Yadav, Nivedita Gupta, et al. (2021) SARS-CoV-2 Spike Mutations, L452R, T478K, E484Q and P681R, in the Second Wave of COVID-19 in Maharashtra, India. Microorganisms 9(7): 1542.
- Zhao LP, Terry Lybrand, Peter Gilbert, Thomas H Payne, ChulWoo Pyo, et al. (2022) Rapidly Identifying New Coronavirus Mutations of Potential Concern in the Omicron Variant Using an Unsupervised Learning Strategy. Res Sq.
- Cool K, Natasha N Gaudreault, Igor Morozov, Jessie D Trujillo, David A Meekins, et al. (2022) Infection and transmission of ancestral SARS-CoV-2 and its alpha variant in pregnant white-tailed deer. Emerg Microbes Infect 11(1): 95-112.
- Fenizia C, Mara Biasin, Irene Cetin, Patrizia Vergani, Davide Mileto, et al. (2020) Analysis of SARS-CoV-2 vertical transmission during pregnancy. Nat Commun 11(1): 5128.
 Mini Review
Mini Review